Introduction
A single night in the sleep lab is absurdly information-dense. We wire up the patient like a low-budget astronaut, record brain activity, breathing, heart rhythm, muscle tone, then boil the whole thing down to a handful of stage labels and an AHI. It’s like running a full-body MRI and only reporting, “Yep, there’s a person in there.”
SleepFM is a serious attempt to stop doing that. It treats clinical polysomnography as a rich, reusable measurement, not a one-off report. Feed it a Polysomnography test and you get a compact representation of the night, an embedding you can reuse to study risk across many outcomes, not just sleep staging.
The bet is simple. If you train a Medical foundation model on hundreds of thousands of hours of PSG, it will learn physiological signatures that correlate with future disease. Not in a mystical way. In the ordinary way biology leaves fingerprints in signals.
Table of Contents
1. Beyond The Hype: What Is SleepFM And Why Does It Matter?

If you’ve seen the social media version, SleepFM looks like a magic trick. One PSG, 130 diseases, done. The reality is more interesting, and more useful.
It’s also not a random preprint doing laps on Twitter. The work landed in Nature Medicine, which usually means reviewers demanded more than vibes.
SleepFM is a multimodal foundation model trained on roughly 585,000 hours of polysomnography from about 65,000 participants across multiple cohorts. It ingests several PSG modalities, including EEG and EOG, ECG, EMG, and respiratory signals. Instead of being hard-coded to one montage, it was built to tolerate the messiness of clinical practice, channels missing, channels reordered, different sensor layouts.
The trick is not “predict everything.” The trick is “learn a good representation.” The model produces latent embeddings that capture the physiological and temporal structure of sleep, then you fine-tune lightweight heads for downstream tasks like disease risk, sleep staging, and apnea classification.
| What Clinicians Usually Get From PSG | What The Model Adds | Where It Fits In Real Workflows |
| Sleep stage report, respiratory events, AHI, oxygen metrics | A learned “night vector” and reusable SleepFM embeddings | Retrospective screening on archived studies, triage flags, research cohorts |
| Rules and thresholds tied to one task | One representation that supports many downstream tasks | Label-efficient modeling, transfer learning to new cohorts |
| A PDF you read once | Features you can reuse as questions change | Continuous improvement as your Sleep dataset grows |
| Costly lab night with limited reuse | Better return on Polysomnography cost | A second layer of value, beyond billing codes |
1.1 What “Foundation Model” Means In This Context
In imaging, a foundation model learns general structure from many scans, then fine-tunes to specific labels. SleepFM does the same for PSG. It learns what a stable night looks like, what fragmentation looks like, what cardiopulmonary stress looks like, and it learns those patterns across cohorts.
This is not a wearable system. It is not built for Apple Watch or Fitbit grade summaries. It is PSG-first. Think “clinical data, clinical signal quality.”
2. The Science: How One Night Of Sleep Predicts Mortality And Disease
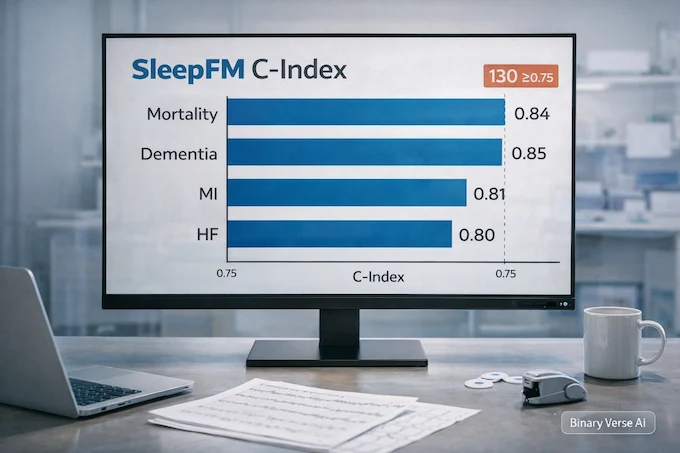
The paper reports SleepFM delivers strong ranking performance across many outcomes, clearing a C-index or AUROC of at least 0.75 for 130 conditions. The headline scores include all-cause mortality around 0.84, dementia around 0.85, myocardial infarction around 0.81, and heart failure around 0.80.
Those numbers matter for one reason. They say the night contains risk signals that demographics miss. Age, sex, and BMI are useful. They are also blunt instruments. A Polysomnography test captures physiology in motion, arousals, autonomic shifts, intermittent hypoxia, and the timing of all of it.
The study also treats outcomes with more care than most “AI predicts disease” claims. Diagnoses are mapped from ICD codes to phecodes, and the timestamp for a condition is set to the earliest matching code. Positive cases are defined so the first diagnosis occurs after the sleep study, not before. AUROC is computed on a multi-year horizon for many conditions. That is the boring work that makes the exciting numbers believable.
2.1 Why C-Index Is The Right Lens For Risk
For disease prediction, you often care about ranking. Who is higher risk than whom. C-index is built for that. It rewards models that assign higher risk to patients who experience events earlier, and it works naturally with censoring. If you’re thinking about screening and follow-up, ranking is usually what you act on first.
3. Prerequisites: Hardware And Data Requirements For Doctors
Let’s get the “no-go” list out of the way. This does not run on smartwatch sleep summaries. It expects clinical PSG signals, the kind you get from a sleep lab, not from a wrist accelerometer.
3.1 System Requirements
The reference implementation is built for Linux. In the released codebase, the authors report testing on NVIDIA GPUs including A100 class hardware. Smaller cards can work by reducing batch size. Plan on at least 32 GB RAM. For preprocessing and training runs that do not feel like punishment, use an 8-core CPU or better.
If you want a mental model for compute, the paper reports that one epoch of large-scale pretraining took on the order of tens of hours on a single A100 class GPU. On a smaller public cohort, you should expect hours, not minutes. Fine-tuning downstream heads is much faster.
3.2 Data Requirements
You need PSG files in .EDF format or an equivalent that you can convert. You also need consistent channel naming and time alignment. PSG is messy, and foundation models tolerate variability, not chaos.
The preprocessing described in the paper resamples signals to 128 Hz and segments them into 5-second windows that become the model’s input tokens. That single detail has downstream implications, every signal must be aligned, and missing segments need consistent handling.
If you are building a Sleep dataset from your clinic archive, standardize three things:
- Channel mapping to modalities, EEG, ECG, EMG, respiratory.
- Time alignment and missing data handling.
- A clean link between PSG IDs and outcomes.
Do those, and SleepFM becomes straightforward to evaluate. Skip them, and you’ll spend your budget debugging filenames.
4. Step-By-Step Guide: How To Set Up SleepFM For Research

This section is the “get it running” path. It is written for a clinical researcher who can run commands, or for a data scientist embedded with a sleep lab.
4.1 Step 1. Create The Environment
git clone https://github.com/zou-group/sleepfm-clinical.git cd sleepfm-clinical conda env create -f env.yml conda activate sleepfm_env
If you prefer pip:
python -m venv .venv source .venv/bin/activate pip install -r requirements.txt
4.2 Step 2. Preprocess EDF Into Model-Ready Tensors
The pipeline expects tensors produced from raw EDF. The repo includes a preprocessing script for EDF conversion.
python preprocessing/preprocessing.py \ --dataset_root /data/mesa \ --output_root /data/mesa_hdf5
The output should mirror your EDF filenames. If your EDF is mesa-sleep-0001.edf, your processed output should keep that identity stable. Every downstream step depends on this being boring and consistent.
4.3 Step 3. Generate Embeddings
Once the data are in the expected format, generate embeddings. Think of embeddings as the distilled summary of the night.
python sleepfm/pipeline/generate_embeddings.py \ --config configs/config_set_transformer_contrastive.yaml \ --checkpoint sleepfm/checkpoints/model_base \ --data_root /data/mesa_hdf5 \ --output_dir /data/mesa_embeddings
Under the hood, the model starts with one-dimensional convolutions for feature extraction, then uses channel-agnostic attention pooling to handle varying channel number and order across cohorts. A transformer encoder captures temporal dependencies over a 5-minute context window. This is the part that turns “signals” into “structure.”
4.4 Step 4. Fine-Tune For Disease Risk With CoxPH
The disease head in the paper uses a Cox proportional hazards style objective. That lets the system learn risk from time-to-event data.
python sleepfm/pipeline/finetune_diagnosis_coxph.py \ --config sleepfm/configs/config_finetune_diagnosis_coxph.yaml \ --checkpoint sleepfm/checkpoints/model_diagnosis \ --embeddings_dir /data/mesa_embeddings \ --labels_path /data/outcomes/cox_labels.csv \ --output_dir /data/runs/diagnosis_coxph
Your cox_labels.csv needs, at minimum:
- patient_id
- time_to_event (or follow-up time)
- event (0 or 1)
- label (phecode, ICD-derived group, or a task-specific outcome)
Start with one outcome. Prove the wiring is correct. Then scale.
4.5 Step 5. Use Embeddings In Plain Python
You don’t need deep learning code to start using embeddings. A first pass can be as simple as loading arrays and fitting a model using standard Python.
from pathlib import Path
import numpy as np
emb_dir = Path("/data/mesa_embeddings")
first = next(emb_dir.glob("*.npy"))
x = np.load(first)
print("Embedding shape:", x.shape)
print("All finite:", np.isfinite(x).all())From there, you can feed embeddings into survival models, tree models, or linear baselines. SleepFM is happiest when you treat it as a feature extractor first, then as a system you fine-tune once you trust the data pipeline.
4.6 Step 6. Sleep Staging And Apnea Tasks
If your goal is sleep staging, the repo includes a fine-tuning and evaluation pipeline.
python sleepfm/pipeline/finetune_sleep_staging.py \ --config configs/config_finetune_sleep_events.yaml python sleepfm/pipeline/evaluate_sleep_staging.py \ --config configs/config_finetune_sleep_events.yaml
For apnea, the same embedding workflow applies, generate embeddings, then train a classifier head. The paper reports strong accuracy for apnea presence and moderate accuracy for severity classification.
5. Case Study Application: Detecting Sleep Apnea Severity
A sleep lab already has a workflow for apnea, manual scoring, event labeling, then review. SleepFM does not replace that. It adds a fast, consistent signal that can triage and sanity-check.
The reported accuracy for apnea presence classification hits 0.87, which is high enough to be useful as a flagging layer. The severity task is harder, the four-class classification accuracy is around 0.69, which still helps as a rough stratifier.
The value is not in the number alone. The value is in what you can do with it:
- Push high-risk studies to the front of the review queue.
- Detect systematic scoring drift across technicians.
- Identify borderline cases where a second look is warranted.
That’s how AI sleep earns trust, not by replacing humans, but by reducing the odds of a missed signal on a busy day.
6. Validating The Results: MESA And SHHS Sleep Datasets
Generalization is the entire game. Many models look great on the cohort they grew up in, then fall apart in the real world, where sensors differ and populations shift.
SleepFM addresses this in two ways. First, the architecture is explicitly channel-agnostic, designed for variability across cohorts. Second, it evaluates transfer learning on SHHS, a large dataset excluded from pretraining. In plain terms, the model is tested on data it has never seen during representation learning.
If you are deciding whether to invest, you should care about this setup more than any single metric. A foundation model is valuable when it transfers, because your clinic is not the training set.
6.1 The Sleep Dataset Reality Check
MESA and MrOS are public and well-studied. SHHS is large and diverse. Your local Sleep dataset will still differ. Patients, comorbidities, scoring rules, hardware, and referral patterns all vary.
So use the public results as a directional signal, then run your own validation. That is how you turn a paper result into a clinical research program.
7. Limitations: What SleepFM Cannot Do Yet
SleepFM is impressive. It also has limits that matter in real deployments.
| Limitation | What It Means In Practice | Practical Mitigation |
| Interpretability is limited | A high risk score does not explain the physiological driver | Pair outputs with modality ablations, review representative traces, add explanation layers |
| Dataset bias toward sleep clinic populations | Predictions may shift in healthier or underserved populations | Recalibrate on local data, stratify by demographics and referral patterns |
| Temporal drift | Performance can degrade as practice patterns and populations change | Monitor calibration, re-train heads periodically, keep an external holdout |
| Task definitions are coarse in some settings | Apnea severity is framed as thresholded classes | Evaluate on your operational labels and consider regression or event-level tasks |
7.1 The Black Box Problem, With A Clinical Accent
Deep representations are hard to explain. That’s not unique to this model, it is a general issue with learned features. The fix is not pretending interpretability is solved. The fix is building layered evidence, ablations by modality, calibration curves, subgroup analyses, and a habit of treating model outputs as prompts for investigation.
8. The Future Of AI Sleep Analysis In Clinical Practice
The most exciting use of SleepFM is not “replace sleep staging.” It is shifting PSG from reactive diagnosis to proactive risk management.
Imagine a near-future workflow:
- Every clinical PSG automatically produces embeddings.
- A downstream model screens for elevated long-term risk, cardiovascular, neurocognitive, renal.
- The EHR gets a quiet flag, not an alarm, a suggestion for follow-up, not a diagnosis.
- A clinician decides what to do with that signal.
That’s the kind of system that could justify the Polysomnography cost to health systems that constantly ask, “Why are we doing this test?”
There’s also an obvious consumer endpoint. An AI sleep coach that is grounded in real physiology, not just bedtime reminders, could combine PSG-derived insights with longitudinal wearable tracking. SleepFM itself is not the wearable model, but it can define the representation that future systems aim to approximate.
9. Conclusion: Is Your Clinic Ready For AI?
SleepFM turns a Polysomnography test into something closer to a reusable biomarker. It does not replace clinical reasoning. It upgrades what you can do with data you already collect.
If you run a sleep lab or a research group, the next step is concrete:
- Choose one archived cohort.
- Standardize the channels and outcomes.
- Generate embeddings.
- Validate risk prediction on your population.
- Decide whether the signal changes decisions.
If you want to stop wasting the richest night of data in medicine, start a pilot with SleepFM on your existing archive. Bring a data scientist into the room, define outcomes like you mean it, and let your Sleep dataset earn its keep.
What Is A Medical Foundation Model?
A medical foundation model is a large AI trained on huge amounts of unlabeled medical data, then adapted to many tasks with minimal extra training. SleepFM is an example, it learns from polysomnography signals and transfers that knowledge to disease risk prediction and sleep staging.
Will Sleep Techs Be Replaced By AI?
Unlikely. SleepFM can speed up scoring and surface risk patterns, but sleep technicians still own data quality, artifact handling, edge cases, and clinical context. Think “second reader at scale,” not a replacement.
What Is A Polysomnography Test For?
A polysomnography test is the gold standard overnight sleep study used to diagnose sleep disorders. It records signals like EEG, ECG, oxygen, airflow, and respiratory effort. SleepFM uses that high-resolution physiology to build AI sleep representations linked to future health risk.
Can AI Predict Sleep Apnea?
Yes. SleepFM can classify sleep apnea presence with about 87% accuracy and estimate severity as well. It learns patterns in airflow, oxygen, and respiratory signals that often take humans longer to piece together across an entire night.
How Much Does A Polysomnography Analysis Cost?
Polysomnography cost varies by country and hospital, but in many clinics it’s often around $1,000 to $3,000 per night. SleepFM does not reduce that bill by itself, but it can increase the value you get from the same expensive test by extracting more clinically relevant signals.

